Abstract
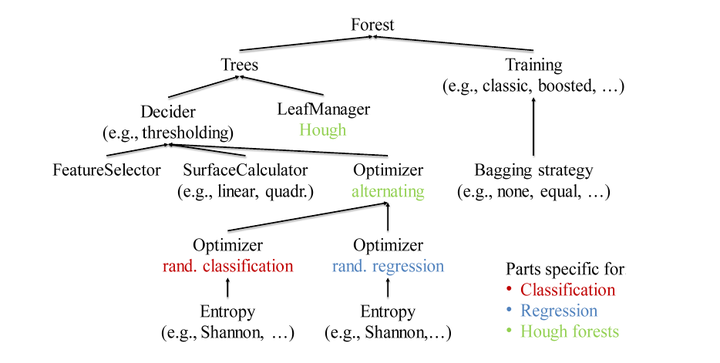
Since the introduction of Random Forests in the 80’s they have been a frequently used statistical tool for a variety of machine learning tasks. Many different training algorithms and model adaptions demonstrate the versatility of the forests. This variety resulted in a fragmentation of research and code, since each adaption requires its own algorithms and representations. In 2011, Criminisi and Shotton developed a unifying Decision Forest model for many tasks. By identifying the reusable parts and specifying clear interfaces, we extend this approach to an object oriented representation and implementation. This has the great advantage that research on specific parts of the Decision Forest model can be done `locally’ by reusing well-tested and high-performance components. Our fertilized forests library is open source and easy to extend. It provides components allowing for parallelization up to node optimization level to exploit modern many core architectures. Additionally, the library provides consistent and easy-to-maintain interfaces to C++, Python and Matlab and offers cross-platform and cross-interface persistence.